Modelling the Brain[]
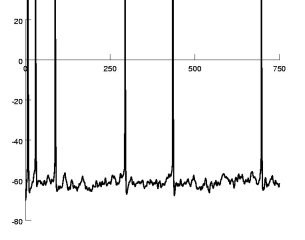
(Electronic Vision(s) Group, Heidelberg)
Nearly all previous theoretical approaches for understanding biological nervous systems were based on models with simplified and homogenised neurons and synapses, simplified connection patterns and simple dynamics converging towards a set of point attractors. It is well known that the collective and complex behavior of large populations of such units may lead to a distributed representation of information.
These now classic holographic theories have recently been the subject of renewed interest addressing the framework of complex dynamical systems, such as the dynamics-based computing, computing using neuronal diversity, or the liquid computing paradigms. From experimental studies, it is becoming increasingly clear that a diversity of functional assemblies are coexisting dynamically in the same anatomical network. The organisation of computations in neural microcircuits can only be understood on the basis of activity-dependent regulation and plasticity mechanisms that optimize such circuits for diverse tasks.
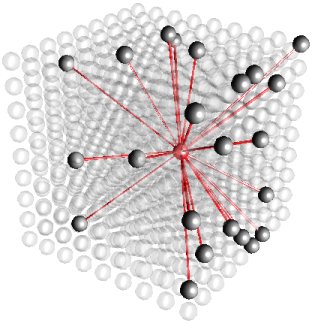
(Electronic Vision(s) Group, Heidelberg)
One reason for the relatively slow progress in the field of theoretical neuroscience is the difficulty of evaluating models with adequate test facilities. The invention of the digital computer has been a great improvement. Today, it is theoretically possible to test any model of neural function if it can be described by a mathematical formalism and if the resulting equations can be solved numerically. While this is in principle true for nearly all existing models, the computing time needed for this calculations becomes the limiting factor. Especially if plasticity, diversity and development are part of the model, the available computing power will be quickly insufficient to explore the timescales involved.
In order to solve the model equations, all digital simulations rely on the repeated execution of simple operations on data stored in some kind of memory. This is fundamentally opposite to the realisation in the human nervous system, where 100 billions of neurons and about 1016 synapses operate in parallel in continuous time. There is an enormous gap between nature and simulation, which reaches a complexity in the order of 103 neurons in real-time with a simple integrate-and-fire model and conductance based synapses on the fastest available microprocessors.